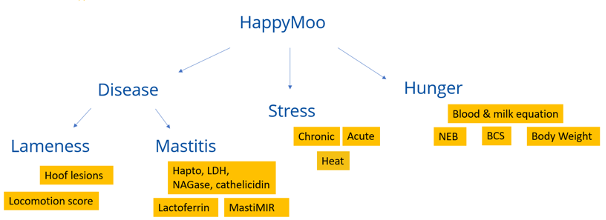
Lameness
Two indicators of lameness were investigated. The first one is the locomotion score. These data came from the D4Dairy project and the others came from Belgium.
Several models have been tested on these data to predict lame cows like the decision tree, the random forest, ... but without very convincing results. Better results were obtained thanks to machine learning methods but the external validation showed that the model could not be used in the field for the moment.
The second indicator investigated for lameness was hoof lesions. The machine learning methods tested by IDELE allowed us to predict healthy animals while we were interested in the opposite.
Mastitis
Four sub-topics were investigated.
- The existing MastiMIR model has been tested in the field during HappyMoo in Germany and France. Microbiological analyses were done to see if bacteria were found in animals that were predicted to have mastitis. It was found that 90% of these animals had different bacteria. Then, the animals with the lowest SCC were also tested to see if bacteria were present or not.
- The second one is the lactoferrin equation. This equation could not be updated before the end of the project because we were still waiting for standardized spectra.
- From historical data, we also had dataset on haptoglobin (marker of inflammatory response in the udder). In the project, we supplemented this data with new samples collected during the project period and among several countries. Several models were tested by ULiège and Seenovia on the data and the best results were obtained using the loglmnet and glmnet methods. An important part of the results obtained are explained by the somatic cells count. They found a strong correlation between haptoglobin and somatic cell count.
- Research has also been done on lactate dehydrogenase (LDH) and NAGase, two enzymes identified as biomarkers of mastitis. The equations were done by CRA-W. The MIR models predicting LDH are not good enough to be used in the field while the results on NAGase are much better.
Hunger
It is probably for this topic that the most indicators have been investigated.
- The negative energy balance was studied through 3 parameters: the energy balance (EB), the dry matter intake (DMI) and the feed efficiency (FE). The models were developed by the LKV-BW and the results obtained allowed a good prediction of these parameters.
- The existing equations predicting milk biomarkers could be updated with the new data. The models built for the indicators BHB, acetone and citrate gave good results and the models could be implemented. Other parameters such as Glucose-6-phosphate, isocitrate and glucose free were also tested but are not yet usable at this time.
- Work on the body condition score (BCS) must continue in order to improve the performance of the developed equation.
- Bodyweight
An external validation with data from restriction feeding trials was done. Two pieces of information emerge:
- Oleic acid is a biomarker that responds very well to restriction.
- EB, DMI and FE are correlated to oleic acid predictions
Stress
The stress working group started from zero, and after a large literature review, an experiment was conducted at CRA-W to identify chronic stress indicators among potential biomarkers.
This study led to the publication of an article available here: https://doi.org/10.1016/j.animal.2022.100502
After identifying hair cortisol and blood fructosamine as chronic stress biomarkers, a large-scale sampling was organized among European countries in order to collect reference values associated with milk MIR spectra. This will allow to study the possibility of milk MIR spectra to gather indication on chronic stress status of dairy cows.
A parallel study was done on acute stress using data from respiration chambers and trimming datasets. The first results were quite interesting and this topic would be interesting to explore in the future.
The stress working group also explored heat stress topic through the development of a model predicting the THI indicator with fair accuracy. In a second step, the University of Liege assessed the relevance of this indicator and several aspects suggests that it would be better than the usual traits (milk, fat, protein). The research is going further to explore how it could be used in routine to evaluate heat stress.
Deliverables
Regarding deliverables, several models have been implemented and are available for milk recording organisations.
These models could be provided in the form of APIs thanks to dockerization learned during the knowledge sharing of our IT teams.
The HappyMoo deliverables available are:
MastiMIR (Germany, France), milk biomarkers model (NAGase, BHB, acetone, citrate in milk), BCS, THI.
The whole partnership hopes to continue working together on these tools. Fingers crossed that we will meet again in 2023.